Research projects Bastians lab
Department of Molecular Oncology
Our lab is focusing on the mechanisms of Chromosomal Instability (CIN) in human cancer. CIN can be categorized into structural CIN (S-CIN) causing structural chromosome aberrations (e.g. deletions, translocations or amplifications) and numerical CIN (W-CIN) leading to aneuploidy. While S-CIN can be due to defects during DNA replication or caused by faulty DNA repair, W-CIN is a direct consequence of chromosome missegregation during mitosis. Intriguingly, S-CIN and- W-CIN are typically detected concomitantly in cancer cells suggesting that the routes to both forms of genomic instability are inter-linked. However, it remains largely unknown how CIN is induced in human cancer and how CIN affects tumor evolution.
Past work from our lab has provided new insights into the mechanisms of numerical CIN that leads to the induction and evolvement of aneuploidy. Mechanistically, we discovered that abnormally increased microtubule dynamics within mitotic spindles can act as a key driver for whole chromosome instability and the induction of aneuploidy. In fact, this type of mitotic defect is highly prevalent in chromosomally unstable cancer cells.
The main focus of our lab projects is to understand how mitotic errors are generated in human cancer cells and how CIN might be suppressed in a therapeutic context.
Projects
Project area 1: Cross-talks of DNA replication stress and mitotic dysfunction to unravel molecular routes to chromosomal instability in human cancer
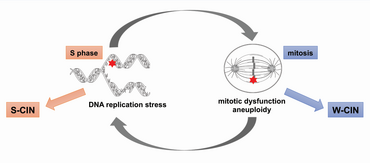
Since S-CIN and W-CIN often occur simultaneously, we are investigating possible mechanisms that link these two forms of CIN.
Interestingly, cancer cells frequently exhibit defects during DNA replication, a condition known as replication stress that can directly trigger S-CIN. Recently, we found that replication stress coincides with perpetual chromosome missegregation and W-CIN, suggesting mechanistic cross-talks between replication stress and mitotic dysfunction and vice versa.
To address these highly relevant cross-talks, our lab has initiated a nation-wide DFG-funded collaborative research unit (FOR2800, www.for2800.de) entitled “Chromosome Instability: cross-talks of DNA replication stress and mitotic dysfunction”. The FOR2800 which combines the expertise from seven german laboratories focusing on mitosis, DNA replication and genome dynamics. As part of our research consortium our lab is addressing the question of how DNA replication stress during S phase of the cell cycle is linked to chromosome missegregation and W-CIN in mitosis. For this, we are investigating various signaling pathways that link DNA replication to mitosis and we are investigating the cancer-relevant causes of DNA replication stress as a trigger for CIN.
Project area 2: Mechanisms of chromosomal instability in pancreatic cancer
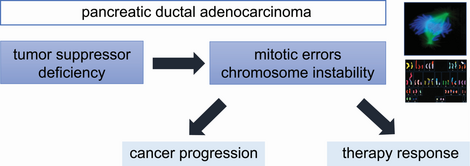
Pancreatic ductal adenocarcinomas (PDAC) belongs to the most aggressive forms of cancer and, accordingly, PDAC shows very high levels of chromosomal instability that drives tumor evolution towards highly aggressive forms of pancreatic tumors. However, the underlying mechanisms contributing to CIN and aggressiveness in PDAC are little understood.
As part of the DFG-funded Clinical Research Unit 5002 (KFO5002, www.gccc.umg.eu/en/cru-5002/) we are investigating the basic mechanisms of CIN in PDACs. Here, we are focusing on tumor suppressor pathways known to be frequently lost in PDACs to unravel the molecular causes for CIN and to investigate the impact of CIN on tumor progression and therapy response in this devastating form of cancer.
Project area 3: The role of tumor suppressors and oncogenic pathways in the regulation of chromosome segregation and chromosomal instability in cancer
Since increased microtubule dynamics in mitosis can act as a highly common trigger for W-CIN, we are investigating the role of tumor suppressors and oncogenic pathways in the regulation of microtubule dynamics and CIN. In fact, we identified already important tumor suppressor suppressors including BRCA1 or TP53/TP73 as being required for proper regulation of spindle dynamics. Also, we found that overexpression of oncogenes including AURKA and CEP72 are sufficient to trigger CIN. We are now aiming to understand the molecular role of these cancer-relevant proteins in the regulation of mitosis and in the maintenance of genome stability.
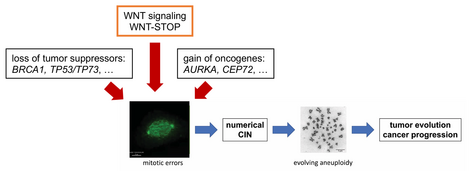
An additional important and highly cancer-relevant signaling pathway, which we found to be involved in the regulation of chromosome segregation, is the Wnt signaling pathway. Interestingly, we identified a requirement of beta-catenin independent Wnt signaling, a pathway now known as Wnt-mediated stabilization of proteins (WNT-STOP), for faithful chromosome segregation.
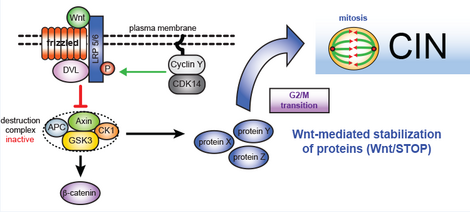
As part of the collaborative research center 1324 (SFB1324, www.sfb1324.de) we are investigating how Wnt signaling and WNT-STOP maintains genome stability by ensuring proper execution of mitosis.